Hemocytometer?
A hemocytometer is a square chamber carved into a piece of thick glass that has a specific depth. It is used to calculate the density of cells in suspensions. Counting cells can’t be done directly from the flask because you don’t have an order of magnitude of the volume of cells you are seeing. You also don’t have a specific area that you can count every single time in the same way, and you don’t know if the portion you have counted is statistically significant or not. What I’m trying to tell you is: there’s a standard way.
It’s called hemocytometer (or hemacytometer / haemocytometer, but not hematocytometer). Hemo, for blood; cyto, for cell; meter, for measuring. So altogether: measuring blood cells.
“Hey but my cells are not blood cells!!”Ok, ok, wait a sec. Now, when this hemocytometer was invented, the guy (Louis-Charles Malassez) was trying to count blood cells. But in the end, the purpose is to count cells that are in suspension, so any cells can be counted really (as long as you suspend them).
How does a hemocytometer work?
This big square has other little squares inside, following the pattern below:
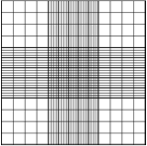

Each of the nine squares inside the big square is equal in surface. In turn, each of the sixteen little squares inside each of the four corner squares is equal in surface. What this tells you is basically, if you count cells in squares that are the same size, you will then be able to take an average of the counts you get, and that will be representative of the cell density of your culture. It is also important to note that if your cell concentration is very high, then you will go and count the cells in the smaller squares rather than the big ones.
“You’re going too fast, I’m lost.”No worries, I didn’t get to the explanation of the protocol yet. Let’s get to the action first.
Counting cells with a hemocytometer
Before I start, I would like to recommend an iPhone app I have been using lately that has made my life a lot easier. It’s called HemocyTap (icon on the right) and it does most of the following itself: it helps you record your data, by acting as a double tally counter, and then performs all the calcs for you and saves the data so that you can review it and even send it to your email!
As I already said, you need to have a suspension sample of your cell culture to perform a cell count. If you are going to determine the viability as well, then you will need to add a viability dye (1:1 dilution usually works best). In any case, this will be your counting solution. Dilute further if your sample is very concentrated. If you need help choosing the basic materials to count cells with a hemocytometer, check our starter kit.
Now that you’re ready, take your hemocytometer, place a glass slide on top (making sure that it does not move; if it does, put some ethanol/water to stick it to the surface) and pipette 10-20μL of you count solution (dilute it if necessary, add viability dyes if you want to tell the difference between live and dead cells). Carefully introduce it in the space between the slide and the hemocytometer (it will go in by capillarity). When the space is filled, you’re done with that side. Repeat with the other big square of the hemocytometer if desired (improved accuracy). Now you’re ready to go to the microscope.
Place the hemocytometer under the microscope, in such a way that you see the first small square of the top big square in the middle of your field of view:
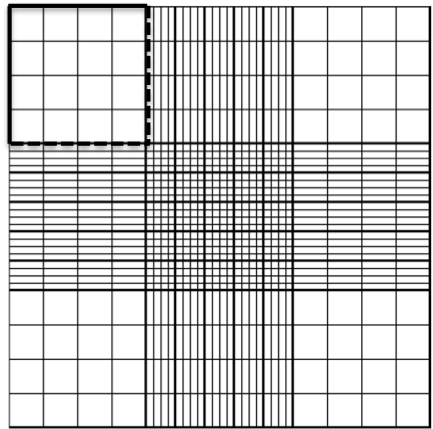
Start counting your cells. Yes, get your tally counters (or iPhone / Android ), feel like those air hostesses checking that everyone’s in, and don’t miss any of your cells! You should decide which two lines of your square you are going to discard. Because you can only count cells once, and some cells will be half-in half-out the square you are counting, it is common practice to choose on which two sides of the square you’ll be counting cells that are touching and on which ones you won’t. But keep these consistently throughout your count. For example:
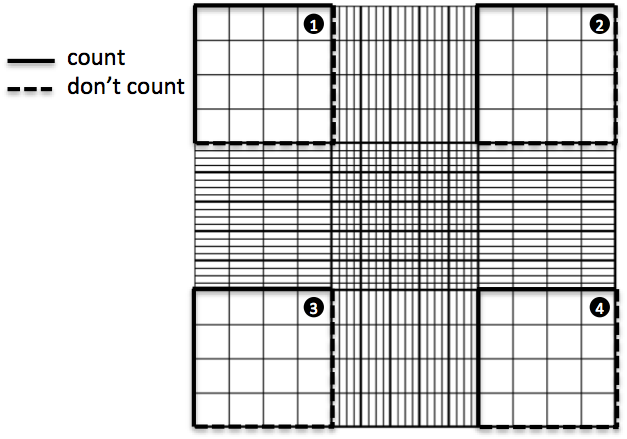
Once you get the cell numbers in the first square, you go to the second (the one on the top right) and do the same thing. Remember, if you counted the cells on the top and right sides but not the ones on the bottom and left sides of the first square, then you must do the same thing with the second one. Keep counting squares until you have enough cells for your count to be statistically significant (big word! well basically, if you did another cell count, the difference between both counts would not be very big). It is recommended that you count at least 100 cells, so that should give you an idea of how many small squares you should count. Keep the counting symmetric (i.e., if you are going to count three small squares per big square, count the top left, bottom right and middle small squares in a big square). Also, if you are not using the app (they are saved for you while you count!), remember to note down your counts every time you finish with a square and reset your tally counter to zero, OR count all the squares and note down the total number, and then also write down how many squares you have counted, so that you can take the average. The advantage of counting all of them is that you don’t need to reset your tally counter to zero every time, but the downside of this is if you are not sure about one of the counts in one square, there is no going back, you’ll have to start again from the first square. This is not the case with HemocyTap, as it keeps the count of the cells in each square without you doing any extra effort of resetting the counters!I definitely recommend using HemocyTap, it saves you many steps and ensures you are not messing it up with the numbers. For a printable protocol, click here. Finally, go to Hemocytometer calculation for a detailed explanation on what to do with your data.

I’m having some problems training some folks here on counting techniques. We work with some very sticky cultures, and I’ve directed them to discard their counts if the left and right sides differ by more than 20%, as an estimate of CoV. Can you direct me to a resource that can do a good job of explaining this method? We will often need to repeat this count multiple times to get numbers that are within 20%, and for the sake of time, I’ve directed them to stop at three repeats, and go back and average all counts. Incorrect statistically, but should be reasonably close.
Hi Stu,
I haven’t come accross this problem so far but I’ve done some research for you:
CV of cell counts according to concentration in 19 cell lines
in here they mention the coefficient of correlation (a good technique might be setting R^2 = 90% and checking if your calculated R^2 for that count falls lower than that), see page 3 table 1
your RHS count falling in the range of the average count in LHS +- 2 times the standard deviation (see equation 1 here
Let me know if that helped!
Cheers
Maria
Hi
I am Vishwas, I am planning to work out a protocol to disassociate the brain tissue and make a cell suspension and stain it with DAPI stain and count the number of neurons to estimate the overall neurons in the brain tissue. There are some publications which came out in the last couple of years, unfortunately there is not enough information to start something from scratch, at least people like me who don’t know much about the haemocytometer.
Currently, I had fixed brain tissue, I mechanically mashed up and centrifuged. But for some reason I dont know how to make up/ determine the intial volume?. I understand that I need to have original volume and concentration, but I dont know how I make up the original volume and concentration. I appreciate if you could help me out, or point me out in the right direction to find a good protocol, which I can follow.
Looking forward to hear from any one of you.
Thank you
Vishwas
Hi Vishwas,
In my understanding, the initial volume is the volume of PBS or counting fluid you resuspend you cells in. Let’s say you have dssociated your cells and you now have a pellet after centrifuging, then just remove the supernatant, add a known volume of PBS or whatever solution is appropriate for your cells, mix and count.
For a video protocol on how to dissociate brain cells, see here.
For a hemocytometer protocol for brain cells, see the last slide of this presentation.
For a protocol on how to use DAPI with brain cells and a fluorescence microscope, see here.
If you combine all three, you should be able to design your own protocol for processing and suspending brain cells, and counting DAPI stained cells with the hemocytometer
Hope that helped! Cheers,
Maria
Thank you Maria for the details, I will try out some of these and let you know how it goes.
No prob! Good luck 🙂
How is it that the solution is diluted with the dye? if the cells are taking in the dye then the dye cannot be the diluent because its volume is being altered.
I was always taught to dilute with saline or phosphate buffer and then add dye. the dye was never used as the diluent.
am I missing something here?
Hi Heather,
You do have to dilute with PBS if your cell density is very high (prior to adding the dye) and then you add the dye. You need to measure how much dye you’re adding, because you’re altering the volume of your counting solution. Unless you’re adding a drop in 1mL+ of counting solution (which would be wasting a lot of cells – hence the use of smaller volumes), the drop in smaller volumes (such as 10uL) will matter and you do have to take that into account. The way you do that is you measure how much dye you add (say, 10uL) and account for the dilution you have created due to the addition of the dye (in this example, 10uL:10uL or 1:1 dilution).
Was that clear? Let me know otherwise what are your specific doubts 🙂
Cheers
Maria
Hello,
I was wondering if you had an explanation for why all nine squares are not counted even though they each have the same volume? I was taught to use the outer four or the central square as outlined on your site, but I know of others who use all nine squares for cell counts. I’m trying to rule out some cell culture issues we have been having and standardize some of our methods. Any input would be greatly appreciated.
Thank you so much!
James
Hi James,
Thanks for your question, it is an interesting one that many readers will find useful!
I think the most important consideration is whether you have enough cells (at least 100 for an estimate) in those 4 corner squares (+ central square) so that the results are significant. Otherwise, you need to dilute your sample less or count more squares (up to all nine of them as your colleagues suggest). If the total count in those 4/5 squares is over 100 cells, in theory there shouldn’t be any difference with the 9 squares counts results. However, the higher the number of cells counted, the more reliable your results and the more likely you are not to find any difference between counting 4/5 or 9 squares. So a good rule of thumb is when the number of cells is under 11 per square, count all nine squares.
I hope that helps, I can do some more research if you tell me the specifics of the differences you’re seeing with your colleagues.
Cheers!
Maria
Hello Maria,
Thank you for answering my question. I have been searching for an answer with mixed results concerning the proper method. What I’m gathering from your response, and the links you graciously provided, is counting a significant number of cells is more important than counting specific squares on the hemocytometer? Because all nine squares have the same area, they should in theory provide the same count as using only the outer four squares provided enough cells are used for the count.
Thanks again!
James
That’s correct James. A minimum of 100 cells per chamber is a good rule of thumb. Although you should always try to keep your counts symmetric in case something went wrong while introducing the suspension and the distribution was not homogeneous (usually this happens when you pipette your cells in too quickly).
The other thing is which squares are easier to count visually. Because the corner and central squares have both vertical and horizontal lines, they are subdivided in more regions than the central top, central left, central bottom and central right. Having more distinct regions helps recognizing areas that have already been counted. And yes, they all have the same area (1mm2).
No prob 🙂
Maria
Hi Maria,
We have used the technique to measure the actual size of a strand of hair,Elodea leaf cell and epithelial cell.While I was searching to write the report I have not acrossed this using.Is it not common?Also,we did this experiment without pipetting any sample.We did not measure the sizes while the hemocytometer was under the microscope together with our samples.First we observed the squares under microscope(magnified at 100x and 400x,respectively) than swicthed the hemocytometer with our samples under ordinary coverslips. Is it a proper way to meaure the actual size of a sample?
Thanks
Hi Asuman,
I do not talk about measuring microscopic sizes here because the most common use of hemocytometers is in calculating cell concentration in a liquid, not measuring cell size (or microscopic measurements of other things like hair). However it does seem other people use it in the way you mention, see for example here. I guess the only thing to take into account for measurement accuracy is how deep the slide and the hemocytometer are comparatively (i.e., if the strand of hair is placed closer to your eye, it is going to appear larger than if it is at the same height as the hemocytometer squares). So the slide where you place the sample should be the same height as the one of the hemocytometer used as your reference. Finally, I would recommend taking pictures and measuring the relative distance in pixels using PhotoShop or ImageJ, in order to convert them back to mm/um.
I hope that was useful, please let me know of any other questions you might have!
Cheers
Maria
I’m wube I have some questions for you: how many squares does one hemocytometer have? 2,why not those squares used for WBC counting is also used for RBC counting and vice versa?
Hi Wube
A hemocytometer has 2 chambers (one at the top and one at the bottom of the slide). Each chamber has 9 large squares, which are subdivided in 16 small squares (for the corner squares) or 25 small squares (for the central squares). Usually the corner squares (and sometimes also the central) are used to count WBC, because they are larger cells. Since RBCs are smaller, they are counted in 5 of the small squares inside the central square, because they are smaller cells. You can see more on which cells are counted where and square sizes in my post: hemocytometer square size.
Hope that was useful!
Maria
Hi Maria,
I am currently doing a lab report on a experiment that I have done on algae involving a hemocytometer. I need a picture of the hemocytometer and I really liked one of the ones that you had on this page explaining what cells to count and what cells not to count. I was wondering, if possible, if I would be able to cite your hemocytometer photograph for my paper?
Thanks,
Elizabeth
Hi Elizabeth,
Of course! It is fine for people to use and cite my images in university reports and in general for work that will not be published or publicly available online elsewhere. Very few of them are from Wikipedia; for those, you will have to credit the authors.
Glad the information found here was useful! Good luck with your report.
Maria
Maria!! you have done a great job! Thanks a ton for making my life easy. stay blessed!
Hi Maria,
I’m doing algae cultivation experiment that I have to count algae concentration by hemocytometer. I have one question to ask you about counting limitation of hemocytometer. Is there information about the lowest cell concentration that can be counted by hemocytmeter?
Thanks,
Thawatchai
Hi Thawatchai,
Any concentration of thefinal solution counted below 200,000 cells/mL (equivalent to 20 cells per corner square counted) would be quite low and result in loss of accuracy.
Hope that helps!
Maria
Hi again Maria. I’ve read several times that it is best to have 100 cells, if I am counting 5 of the 0.2mm squares does it mean there should be 100 in each of the 5 squares or 100 total across all 5? I know this maybe subjective but approximately how much more under or over should we be at? Also if there is too many or too less cells do we continue to dilute until we have near or over 100 then include the dilution factor in our formula?
Hi Plutarco,
There should be 100 cells total across all 5 x 0.2mm squares counted. I would say 10% below or above is ok. Going well below 100 can have an impact in statistical significance of your results, while going well above 100 can make it difficult to count cells accurately (some may overlap, and tracking which ones you have and haven’t counted can become trickier). Hope that helps!
Saludos,
Maria
Hi Maria,
First, thank you so much for putting this information up! It was very helpful.
I have a very basic question to ask that I can’t seem to find the answer to. When you have too many cells in your 1:1 cells/trypan blue mix, do you dilute that mix and if so with what? More trypan blue?
OR do you dilute your total volume of cells and do the whole thing over again?
Bobby
Hi Bobby,
No worries, and glad you find it useful!
It is preferrable to do the whole thing again. However, if you are in a hurry or have a limited amount of cells available for counting, and need to use the sample you have prepared already, I would dilute with PBS or whatever you have used to dilute in the first place.
Cheers
Maria
I’m following tnis up for a student currently studying IB Biology for her Extended Essay
I’m having issues identifying the etchings on the hemocytometer slide under the microscope.
Do you have to stain the slide BEFORE you add the suspension to the slide, or does the colouration of the suspension (after adding the stain) allow you to see the etchings?
Bruce
Hi Bruce,
You usually stain the suspension itself, and that allows you to distinguish which cells are dead (by their coloration). I don’t think the stain will have a significant impact on the hemocytometer. If you are having trouble identifying the lines, I would suggest getting a higher quality hemocytometer, these have lines that don’t fade or are difficult to see.
Maybe I am missing something here, in that case let me know. Good luck!
Maria
Thanks for your reply!
You have confirmed what I had thought. Unfortunately schools sometimes choose the cheaper option, but obviously sacrifice on quality!
It’s great that as a specialist in an area, you make yourself so accessible.
Cheers,
No worries at all! Glad it was useful.
We recently bought the Countess automated cell counter in our lab and we are having trouble using it. After some research, we tried to define the profile settings for counting some cell lines. We mainly count k562, WBC and MSC. I’m doing some comparison with the Neubauer but while doing statistics should I consider +20%/-20% variation acceptable on results from different methods? I’m confused about it. I’ve found little material about the instrument, and we are trying to prove that it’s not working (and seek help from the seller). I’ve seen that over changing the concentration, the counting gets different aswell… Thank you, best regards from Brazil!
Hi Raul,
It can be a bit tricky to get the right settings for each cell line. For common cells like blood cells, there should be information out there on the right settings, see this protocol here.
For each cell counter model, companies publish the results of their studies against hemocytometer counts – you can find the one for the Countess here. As you can see in Fig 3, the difference between hemocytometer counts and automated counts is around 20%. As for hemocytometer variability (by itself), you can see in this other study that it is around 10%, and that also includes the inter-operator variability. However, I would trust more any studies that compare results across different cell counter models such as this one. They report consistently lower counts with the Countess than with the hemocytometer (some reach the 20% difference, see table 3).
With regards to the concentration, hemocytometer counts get less accurate as the sample loaded in the chamber is more concentrated (there are too many cells to identify which ones have been counted and which ones haven’t so in the end some are counted twice and some are not counted). That’s why we usually dilute to get to a concentration that is suitable for visual counting. Machines don’t have that problem, so higher concentrations can be counted.
Let me know if that clarifies a bit, it must be frustrating not to be able to rely on the new machine, but hopefully you either get it working or get a solution from the provider!
Best regards from London!
Maria
Hi,
I am currently trying to count the number of yeast cells alive before I expose them to UV light and then count again using a hemocytometer, except to see how many have died from the exposure. I added the dye (methylene blue) to the solution before I exposed it to UV light to counted the number of cells using the hemocytometer, however they are all blue? What should I do about this as this indicates that none of them are viable before the exposure?
Thanks,
Katie
Hi Katie,
When cells uptake trypan blue, their membrane is not healthy. If you prepared the trypan blue solution correctly, then there must be some issue with your cells.
Otherwise, it could be that your trypan blue solution was not correctly prepared. Note that it should be diluted with isotonic solution (i.e. NOT water), otherwise it will lyse the cells. You can find more information about that here.
Let me know if that helped! Or give me more info on how you prepare the solution so I can try to find out the problem.
Cheers,
Maria
Hi,
I recently try to find the optimal range of cell counting. According to the small cells, I only counted at the central area, 25 boxes (both side of chamber). As recommendation, 100 cells/ chamber. Does it mean there should be 100 in central area of each side or it should be 100 cells in 9 squares? Because some reference recommend it should be 20-50 cells/square. Is there any formula to calculate it?
Moreover, what is the acceptable error of cell counting?
Thanks,
Roch
Hi Roch,
It means 100 cells in the central area of each chamber. If you were counting on the large corner squares, it would be 20-50 cells per square, maybe that’s what the reference you found referred to. You always need to count about 100 cells per chamber, so it’s equivalent between the corner squares and the central square.
With manual counting, there’s always a 10-20% error from count to count (i.e. if you prepare a count from the same sample again and count it yourself) and from operator to operator (if someone else counts the same sample as you).
Hope that helps!
Cheers,
Maria
Hello Maria,
I can’t seem to find an answer to a basic question. What are the dimension the hemocytometer chamber? By chamber I don’t mean the just the grid area, but rather the dimensions, or volume to fill the entire well area of the hemocytometer. I know chamber under the grid has a volume of 0.9 cubic millimeters (i.e. 9 ul). I haven’t yet found anyone to answer my simple question.
Sorry for the late reply K Lai. Were you trying to determine the optimal volume for loading the chamber? I don’t know if anything beyond the grid is entirely standardized among models, because I don’t think it matters generally, as long as capillary action saturates the space.
Dear Maria,
I have been trying to improve my skills in counting cells manually with a hemocytometer. I am usually counting cells in 4 corner squares (top and bottom left and right). I would like to clarify that I need to have minimum 100 cells in those 4 squares, right? What would be the maximum number, then, before I need to dilute the sample?
Also, what would be the appropriate difference of cell count between two chambers?
I keep getting a bigger % error among counts of the same sample. I was wondering if you could recommend some practical tips on how to load the sample to decrease the variability between chambers as well as between counts of the same sample, please
Thank you!
Best regards,
Tim
Sorry this comes so late for you, but hopefully it will help someone else if not you!
The short answer: As many as you can reasonably count, for the uncertainty you desire!
If your cells start clumping up because they’re so dense, probably better to dilute them for a more accurate read.
And if you only need a precision of 3 significant digits, then the high 2-digits per square are just fine and counting more would use more time without giving you more information.
Differences in cell count is most likely due to mixing and pipetting variation. Make sure you mix the solution thoroughly just before taking your aliquot out for the chamber. Unfortunately, there will inevitably be sampling errors when taking such a small sample from a large volume, especially when the cells like to sink and clump.
Diluting the sample could help increase the uniformity, as a larger and presumably better sample of the original is being taken before being subsampled for the hemocytometer chamber.
-Nick
Hello! I am a high school student and, with my group mates, conducting a study which will require us to use a counting chamber for counting and identification of phytoplankton cells. Initially, we had planned to use a Sedgewick-Rafter chamber but due to this material being unavailable to us, we switched to what is available–the hemocytometer. My question then is whether a hemocytometer is appropriate for phytoplankton cells?
Apologies for the late reply. It appears that a Sedgewick-Rafter chamber has a 1-mm thickness vs. 0.1-mm thickness on a standard hemocytometer, and the former has a much larger area for sample. I suspect that the difference may be due to needing to observe actively swimming organisms, like those that may be found in drinking water, and the need to prove levels of microorganism to very low concentration limits.
As long as you can see the cells and there are enough of them, and they’re staying put, a hemocytometer should work well for you.
I am having so much trouble counting cells, and I don’t know what I am doing wrong.
I clean and dry the hemocytometer, then add cell mixture to it using capillary action.
I always mix my cells thoroughly before adding them, but each time there are hardly any cells on the hemocytometer. Even if the cells are very confluent! I know there are more cells in the suspension, but the count is very low (less than 100 cells in the center square).
Instead of pipetting 10uL into the hemocytometer, my PI suggests using a glass pipette to touch the cells and using capillary action to fill the hemocytometer. Could this be the problem?
I am at a loss here because even with high cell density to start with, my counts are low.
100 cells in a big square isn’t that unusual, depending on your cell type, culture dish surface area, and what volume you’re diluting the cells into. 100 cells in one big square is 10^6 cells/mL.
I’m assuming you trypsinize and then pellet the cells, then resuspend in some volume? What are you doing there?
When you load the hemocytometer, do you pipet directly onto the grid and then place the cover slip, or do you pipet into the notch, with the cover slip in place, and allow capillary action to draw it over the grid?
I am Just wondering if you can help me, I am not even sure the comments might even be looked at anymore but i am hoping for the best I am doing a microbiology book for my A levels, and I have looked everywhere but I am a little confused just wonder if you could help me with the question as I have googled it and you can imagine google and it has to be in 30 words, Explain what approach could be used for counting cells that on the border line of a counting square in a haemocytometer to ensure the counting is accurate
Here are some hints: The square defines a specific volume. If you count cells that hang over all the edges, the square you’re counting in is actually bigger than you think it is. If you have two squares next to each other, how do you decide whether a cell hanging over an edge is in one square or another.
I am thankful if you send me a new references about Hemocytometer